About the Folding@home Project
In the growing field of computational biology, medical researchers run complex molecular simulations to help validate therapeutic pharmaceuticals, and develop new ones.
Folding@home is one such project, which began at Stanford University in late October 2000. It’s now based at Washington University in its St. Louis School of Medicine, has additional management from Temple University, and is working on molecular simulation projects from researchers all around the world.
Folding@home runs atomic-level molecular modeling simulations to identify the structure and function of proteins and molecules. These simulations give medical researchers a better understanding of diseases, so they can then make pharmaceuticals which prevent diseases from functioning, evading the immune system, and spreading in the body.
The simulation work Folding@home does is very processor intensive. That’s not a problem, since Folding@home is a massive distributed computing project — it harnesses the spare processing power of volunteers’ computers all around the world to run its simulations. By sending parts of a protein or molecular simulation out to volunteers’ computers for processing and then returning the results, Folding@home has in-effect the fastest supercomputer on Earth.
Coronavirus (COVID-19) was just added to the list of diseases that Folding@home is working on: these include Alzheimer’s, Cancer, Huntington’s, and Parkinson’s.
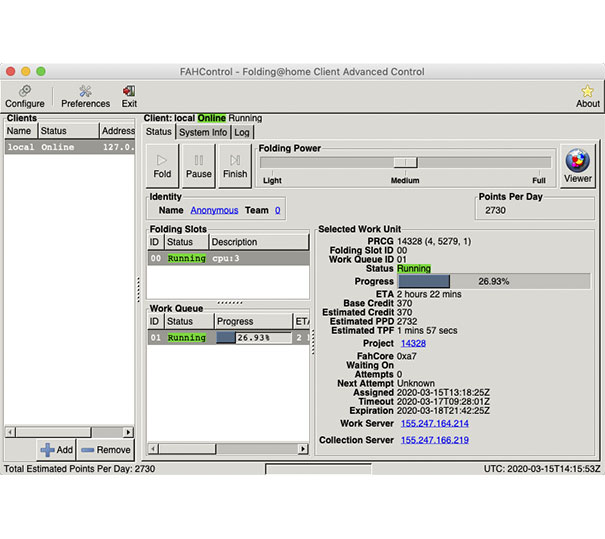
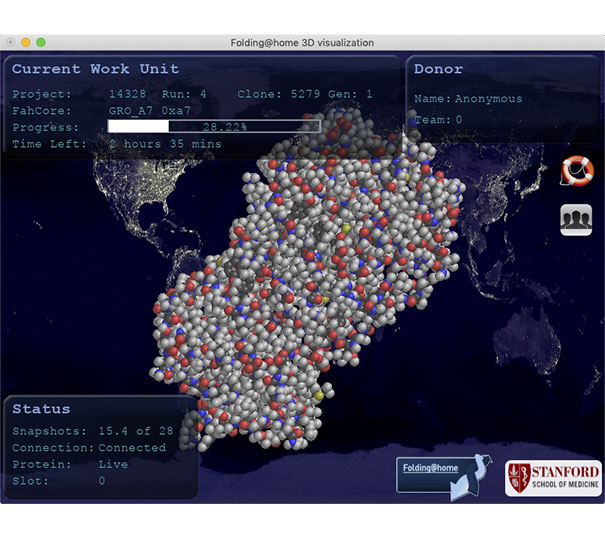
Volunteers can download a free “client program” for Mac, Linux, and Windows, install it, then leave it running in the background while they work on other tasks. The Folding@home client program will download and work on a protein or molecular problem. Completed results are then returned to Folding@home, which will combine calculation results from other volunteers’ computers to form a solution to the protein or molecular problem. Volunteers’ computers are then sent the next protein or molecular problem to work through.
How to Run Folding@home on your Mac
The basics of how to set up the Folding@home client program are explained in a MacRumors story (its easy,) and Windows and Linux versions work in a similar way.
As MacRumors explains, be sure the “Cause” Preference is set to “Any” in order to receive Coronavirus tasks to process (which is the default setting.) Please keep in mind that the “Any” setting will also assign other “high-priority” projects to client programs in addition to Coronavirus-related ones.
Please note that the Folding@home client does not use the Mac’s Graphical Processor Unit (GPU) because of processor library incompatibility reasons. The Mac’s standard Central Processing Unit (CPU) will be used instead.
So many people have joined in Folding@home, that the project’s servers are struggling to keep up. Keep your Folding@home client running, and it will eventually be sent more work to calculate through.
More Information
- Information about Folding@home from Wikipeadia
- Folding@home Download Page
- Folding@home Support page – See the left-column for Installation Guides.
- Folding@home FAQ
- Running Folding@home FAQ
- The Folding@home client program notes what project is worked on with a project number — see the Folding@home Project Page to identify the corresponding project, and read more information about it.
- Folding@home also has an optional “game” aspect to it: you can join teams, and earn points. Read more on the Folding@home Stats, Teams, and Usernames page.PMUG now has a team for Folding@home. If you wish to join us, PMUG’s Team Number is 261185.
In the Web-based controller (client.foldingathome.org) click the “Change Identity” link in the top-left, then type 261185 in the “Team Number” field. In the FAHControl program (the advanced controller,) click the “Configure” button in the top left, then type 261185 in the “Team Number” field. Either way when you’re done, click the “Save” button.
The Name field can be whatever you want: it helps distinguish between computers if you use multiple ones. It can be what you’ve named your computer, or even your alias.
The Passkey field is optional, helps you earn bonus points for completing work early, and helps secure your points. Use the same Passkey if you have multiple computers.
- Folding@home works OK with its default settings, but the Configuration Guide page explains how to customize them.
- Get the latest news from the Folding@home News page and Folding@home Twitter page.
- 5/11/20 – Added new intro paragraphs to explain how computers help in medical research, and how computers help Folding@home.
- 4/11/20 – PMUG Team Info added